Introduction
ABCGrid stands for Application of Bioinformatics Computing Grid.It was designed for small biology laboratories to use large amount of heterogeneous computing resources and access many bioinformatics applications from one master node without being aware of where the data and computational resources are located. ABCGrid is very easy to install, maintain and extend at the premise of high performance. Currently, ABCGrid integrates NCBI_BLAST, Hmmpfam and CE, running on a number of computing platforms including UNIX/Linux, Windows and Mac OS X.
ABCGrid is a server/client application consists of a Master application (ABMaster), a Worker application(ABCWorker) and a User application (ABCuser). ABCGrid is programmed using Java SDK 5.0. Java Runtime Environment (JRE) 5.0 or above(or compatible runtime) is required to run ABCGrid.
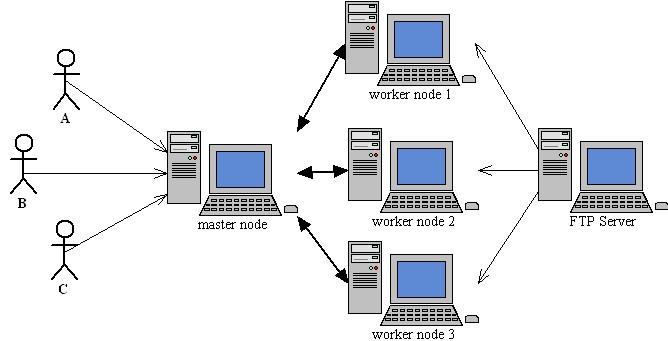
In an typical deployment, you should install ABCMaster in a powerful machine with UNIX/Linux installed. This machine serves as the master node. Every user of ABCGrid should install ABCUser under his own working directory in master node. ABCWorker could be installed in any machines which could be used to do the real computation.
Features
Easy to install, use, manage and extend
Online update for applications and databases installed on worker nodes
Good performance in heterogenenous environment which consist of Windows-PC and powerful UNIX server
Support NCBI_BLAST (blastall, rpsblast,megablast,bl2seq), HMMER (hmmpfam) and CE in current version.
Running on almost all popular platforms include Windows, UNIX/Linux and Mac OS X
Documents
Download
Version:0.1 Date:Dec-25-2006
Source code
Binary package
Source code is released under the GNU General Public License(GPL).
If you want to build ABCGrid from source, you must have JDK 5.0 and Apache-ant preinstalled.
Binary package is a compiled version of ABCGrid for all platforms.
Copyright
2006 Center for Bioinformatics, Peking University.Reference
Ying Sun, Shuqi Zhao, Huashan Yu, Ge Gao and Jingchu Luo. (2007) ABCGrid: Application for Bioinformatics Computing Grid. Bioinformatics
Abstract